Numbers and other invalid (non-IUPAC) DNA characters will be automatically removed. Enter a 20nt ('5-'3) DNA character string representing the used sgRNA guide sequence immediately upstream of the PAM sequence (PAM not included).Tools for Splitting, Applying and Combining Data. The Lawson-Hanson algorithm for non-negative least squares (NNLS). sangerseqR: Tools for Sanger Sequencing Data in R. Biostrings: String objects representingīiological sequences, and matching algorithms. R: A language and environment for statistical computing. This web tool was developed by Eva Brinkman, Christ Leemans and Bas van Steensel from the Bas van Steensel lab.įor more information and to report bugs, please contact Core Team (2013). Stichting het Nederlands Kanker Instituut - Antoni van Leeuwenhoek ziekenhuis (The Netherlands Cancer Institute). Support is given by Data Curators VOF, see our.
#Sequencing chromatogram viewer code
#Sequencing chromatogram viewer software
If you use this software for data analysis in a publication, please cite.The TIDER software is being provided as a free web service for research, educational, instructional and non-commercial purposes only.The output of TIDER is a comprehensive profile of all insertions and deletions (indels) in the edited sample and the frequency of the desired reference (or template) sequence.

The input to TIDER is Sanger sequencing data. It quantifies the efficiency of homology-directed repair ("HDR") in an edited sample by decomposing the sequence trace data from three simple Sanger reactions.
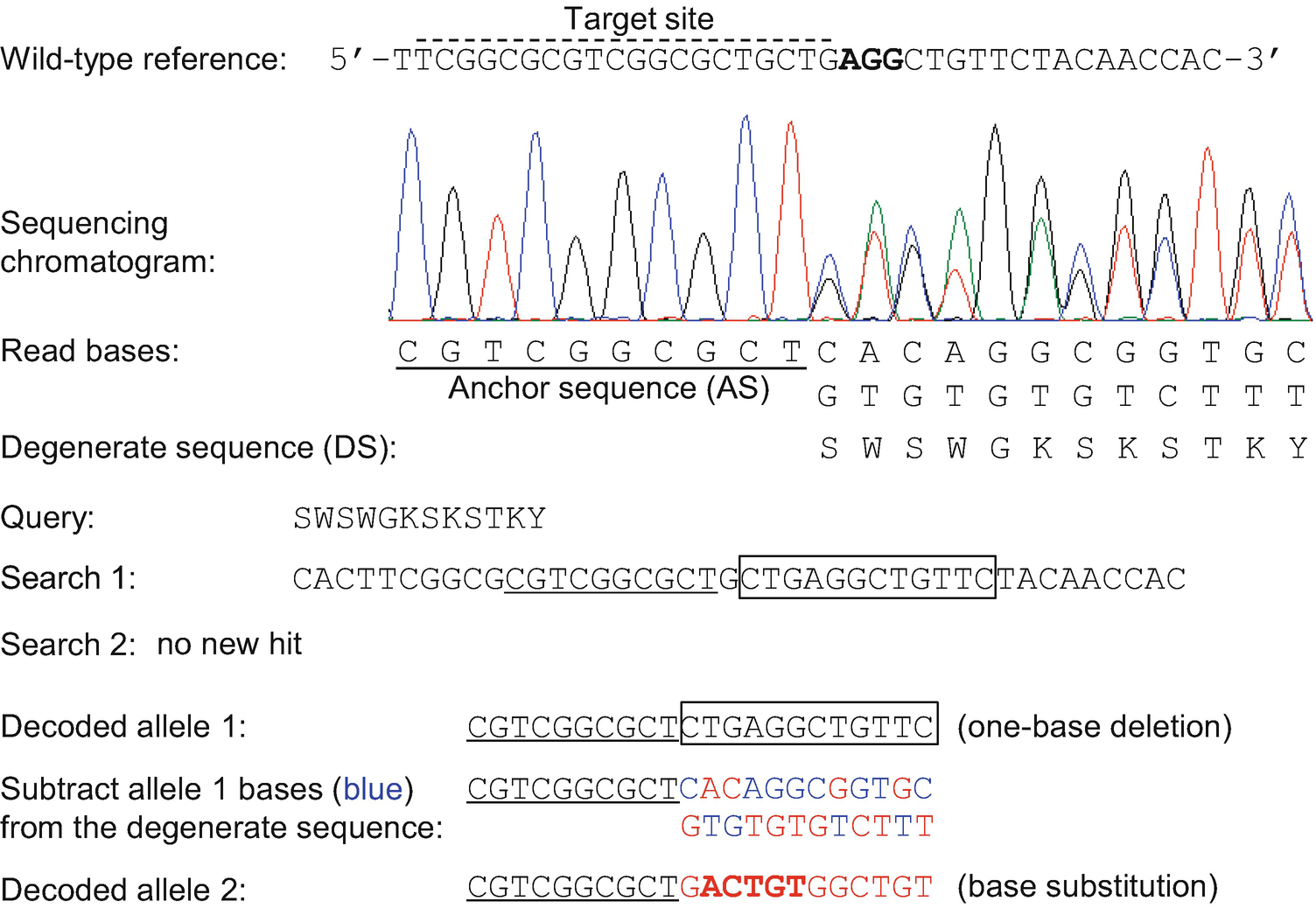
TIDER provides rapid and reliable assessment of template-mediate genome editing. Reference (please cite!): Brinkman et al, Nucl.Drawbacks: Requires some additional wet-lab work to generate a template reference sequence (see Protocol tab).What it needs: Three standard capillary sequencing reactions.For non-templated CRISPR/Cas9, use the original TIDE or TIDE batch web tools. When to use: Quantification of oligonucleotide-templated point mutations and small indels.It also determines the frequency of non-templated indels. What it does: Estimates the frequency of designed (templated) small mutations in a pool of cells transfected with.

TIDER: Easy quantification of template-directed CRISPR/Cas9 editing Purpose
